Published 14 December 2022 in Journal of General Physiology (doi 10.1085/jgp.202213214):
An α–π transition in S6 shapes the conformational cycle of the bacterial sodium channel NavAb
Koushik Choudhury, Rebecca J Howard, Lucie Delemotte
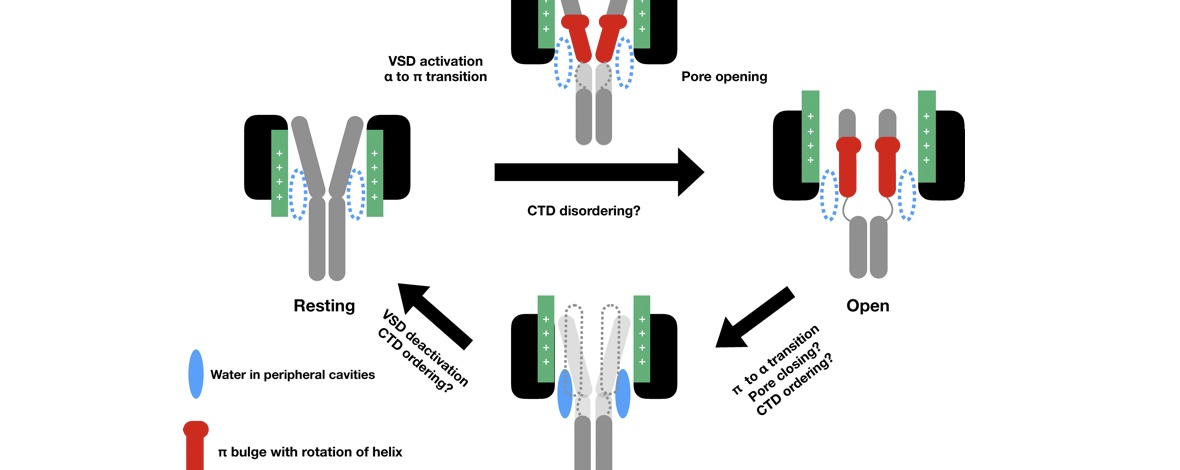
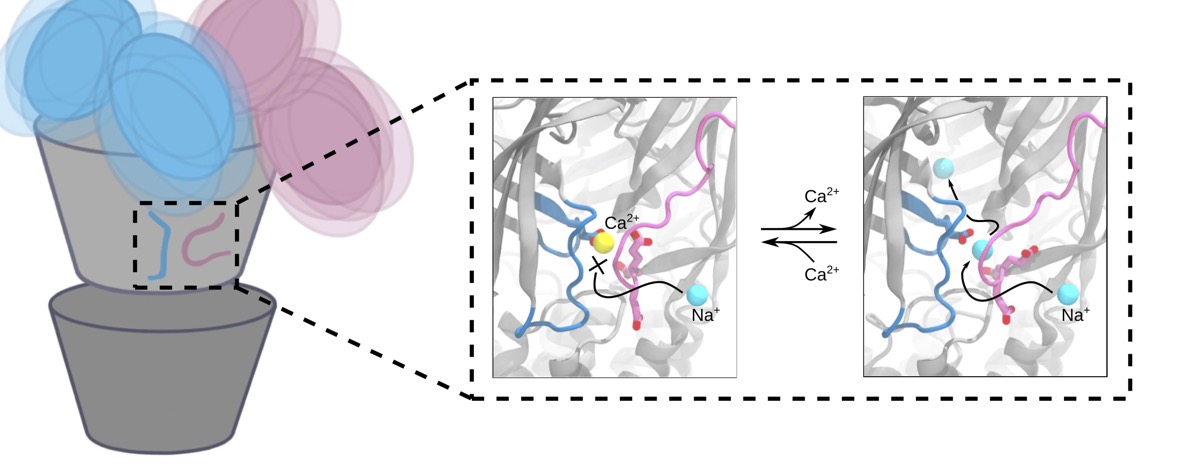
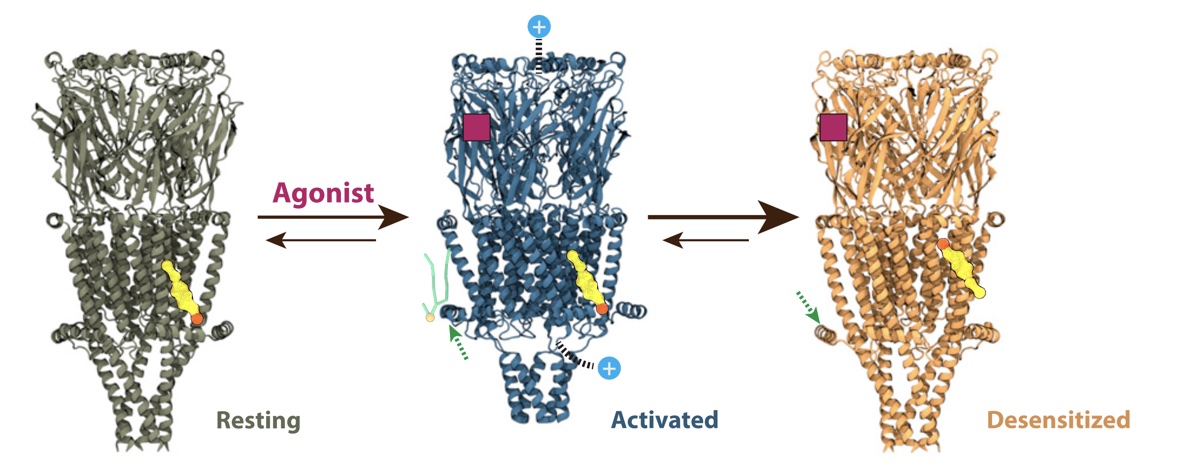
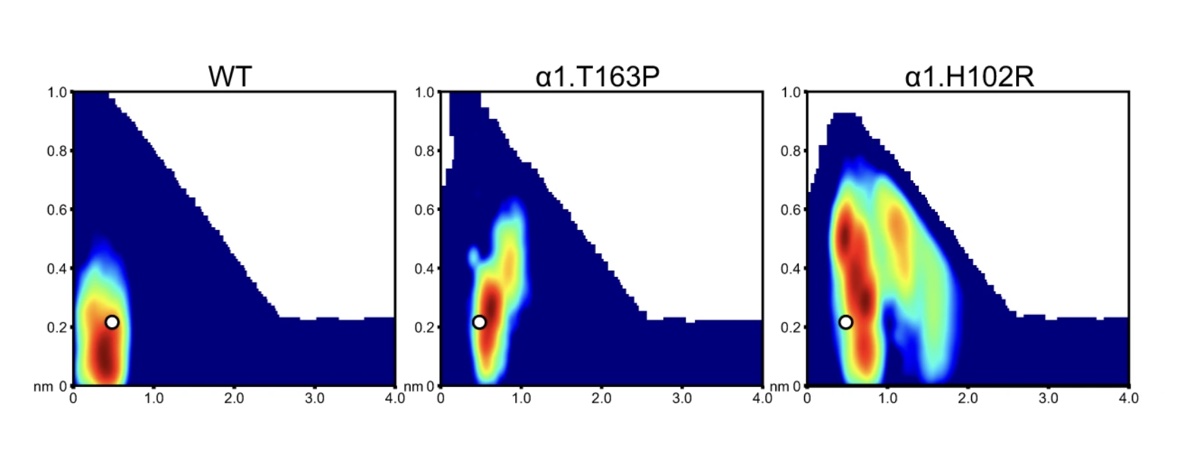
Voltage-gated sodium channels play an important role in electrical signaling in excitable cells. In response to changes in membrane potential, they cycle between nonconducting and conducting conformations. With recent advances in structural biology, structures of sodium channels have been captured in several distinct conformations, which are thought to represent different functional states. However, it has been difficult to capture the intrinsically transient open state. We recently showed that a proposed open state of the bacterial sodium channel NavMs was not conductive and that a conformational change involving a transition to a π-helix in the pore-lining S6 helix converted this structure into a conducting state. However, the relevance of this structural feature in other sodium channels, and its implications for the broader gating cycle, remained unclear. Here, we propose a comparable open state of another class of bacterial channel from Aliarcobacter butzleri (NavAb) with characteristic pore hydration, ion permeation, and drug binding properties. Furthermore, we show that a π-helix transition can lead to pore opening and that such a conformational change blocks fenestrations in the inner helix bundle. We also discover that a region in the C-terminal domain can undergo a disordering transition proposed to be important for pore opening. These results support a role for a π-helix transition in the opening of NavAb, enabling new proposals for the structural annotation and drug modulation mechanisms in this important sodium channel model.
Read the full publication here.